Phot-1 Expression Throughout the Plant
where is it?
As mentioned on the previous page, gene expression may vary in different cell types, tissues, and organs of a multicellular organism like Arabidopsis. Several experiments can be done to show the localization of specific proteins within an organism. A popular technique is to make a fusion protein by adding the gene for Green Fluorescent Protein (GFP) to the end of the gene of interest. In effect, this makes the protein fluorescent. When shone with UV light and viewed under a microscope, the protein lights up wherever it is located. These experiments can also be quantitative: by measuring the amount of light produced at different places or under differing conditions, one can determine the level of expression of the protein of interest.
The University of Toronto has made available a tool which simulates such GFP experiments. It is therefore called the electronic Fluorescent Protein Browser (eFP Browser). Like the Epigenetics Browser, this data is collected from bioinformatic databases. In this case, gene expression is measured through the use of a gene chip. This technology involves a substrate, on which is printed an array of single-stranded DNA probes. A tissue's mRNA is extracted and converted to cDNA via reverse transcriptase, explained further on the BLAST page. These cDNAs are then labelled and exposed to the chip, where they bind to the probes. The more mRNA present in the sample, the greater the signal will be.
The disadvantage of using a gene chip to measure protein abundance is that it indirectly measures mRNA abundance. Some mRNAs may be more highly translated than others, leading to a higher abundance of one protein, even though the gene chip would indicate that they were expressed equally. The main advantages to using gene chips is that they provide a huge amount of information, and they are standardized. A single gene chip experiment can show the expression levels of every single protein-encoding mRNA in an organism. The standardization of gene chips means that the experiments can be repeated and compared. This is exploited in the the eFP Browser, which has data for every Arabidopsis mRNA for various plant organs and tissues. Such genome-wide experiments would be extremely laborious, perhaps even impossible, if done by directly measuring protein abundance with actual GFP fusion experiments.
The University of Toronto has made available a tool which simulates such GFP experiments. It is therefore called the electronic Fluorescent Protein Browser (eFP Browser). Like the Epigenetics Browser, this data is collected from bioinformatic databases. In this case, gene expression is measured through the use of a gene chip. This technology involves a substrate, on which is printed an array of single-stranded DNA probes. A tissue's mRNA is extracted and converted to cDNA via reverse transcriptase, explained further on the BLAST page. These cDNAs are then labelled and exposed to the chip, where they bind to the probes. The more mRNA present in the sample, the greater the signal will be.
The disadvantage of using a gene chip to measure protein abundance is that it indirectly measures mRNA abundance. Some mRNAs may be more highly translated than others, leading to a higher abundance of one protein, even though the gene chip would indicate that they were expressed equally. The main advantages to using gene chips is that they provide a huge amount of information, and they are standardized. A single gene chip experiment can show the expression levels of every single protein-encoding mRNA in an organism. The standardization of gene chips means that the experiments can be repeated and compared. This is exploited in the the eFP Browser, which has data for every Arabidopsis mRNA for various plant organs and tissues. Such genome-wide experiments would be extremely laborious, perhaps even impossible, if done by directly measuring protein abundance with actual GFP fusion experiments.
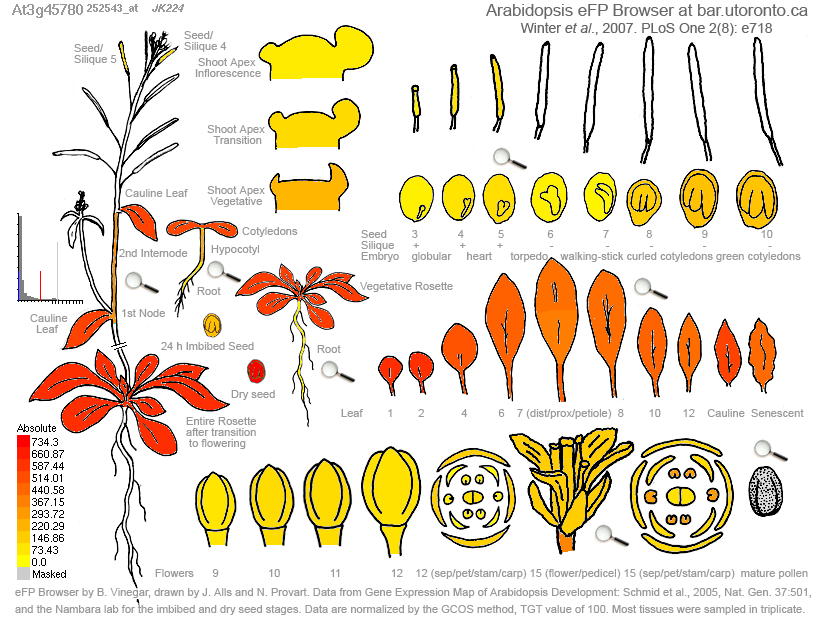
Figure 1. Expression of Phot-1 (At3g45780) in various plant tissues. Highest expression, in red, is shown in the leaves. High levels of expression are also seen in dry seeds. Medium-to-low levels of expression is exhibited in immature floral tissue, mature flower parts, roots, and the embryo. There appear to be no tissues in which Phot-1 is not expressed at all.
Figure 1 above shows the expression levels of Phot-1 (At3g45780) in various tissues and organs and at various developmental stages. Phot-1 appears to be almost ubiquitous throughout the organism. The Epigenetics data, which used immature floral tissue, showed little to no silencing methylation in Phot-1. This is supported by this eFP data, which indicates that Phot-1 is present in moderate levels in this tissue.
Phot-1 Localization in the Root
Of particular interest in regards to the Root Phototropism Review by Kutschera & Briggs (see Overview page) is the location of Phot-1 in the root. This can be inferred through mRNA expression levels, shown in Figure 2 below. In agreement with Kutschera & Briggs proposed model, Phot-1 is located in the growing part of the root, above the calyptra (the root cap). With Phot-1 in this area of the root, light can sensed so that the root can grow in response. Often, but not always, roots respond negatively to light, growing away from it so that they grow away from the surface and down into the soil. High Phot-1 mRNA levels in Figure 2A may represent the "eyes" of growing roots, with Phot-1 acting as a light receptor.
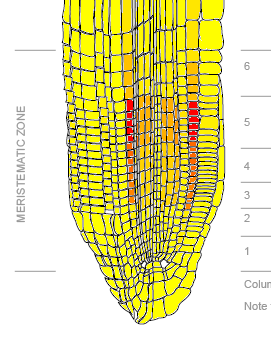
Figure 2A: Phot-1 mRNA expression levels in various root tissues. Yellow represents low expression, orange represents medium expression, and red represents high expression.
Figure 2B-D: Phot-1 mRNA expression by root tissue type. Numbers correspond to the depth of the root, as shown on the right side of 2A. Overall expression is low except in a few tissue types. Expression is highest in the cortex, which is largely composed of undifferentiated cells. Relatively high expression is also seen in phloem companion cells at levels 3-5 (in Figure 2C), phloem pole pericycle at levels 3-5 (Figure 2D), and xylem pole pericycle at levels 3-5 (Figure 2D).
Figure 2B-D: Phot-1 mRNA expression by root tissue type. Numbers correspond to the depth of the root, as shown on the right side of 2A. Overall expression is low except in a few tissue types. Expression is highest in the cortex, which is largely composed of undifferentiated cells. Relatively high expression is also seen in phloem companion cells at levels 3-5 (in Figure 2C), phloem pole pericycle at levels 3-5 (Figure 2D), and xylem pole pericycle at levels 3-5 (Figure 2D).
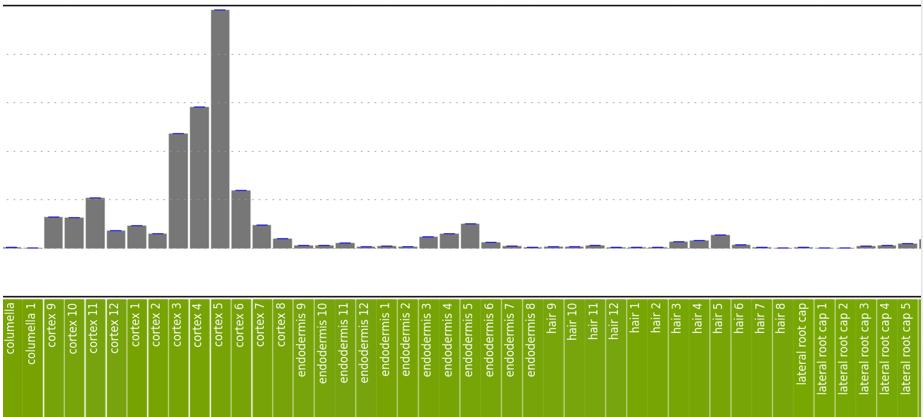
Figure 2B. See figure legend above.
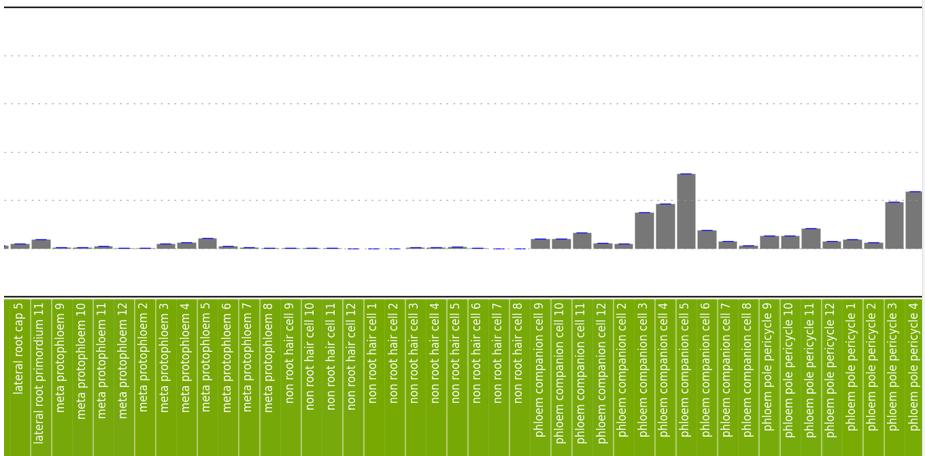
Figure 2C. See figure legend above.
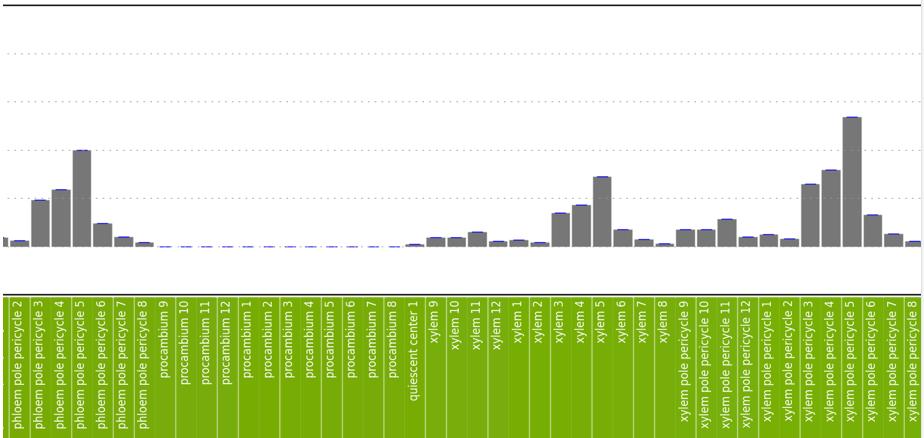
Figure 2D. See figure legend above.
Phot-1 vs Phot-2
The eFP Browser also allows for comparisons in expression levels between two different genes. Relevant here is the comparison between Phot-1 (At3g45780) and the related Phot-2 (At5g58140). Figure 3 below shows the relative expression between Phot-1 and Phot-2 throughout the Arabidopsis plant. Perhaps due to their similar functions, they are distributed roughly equally throughout the plant, though certain differences can be discerned. For example, Phot-1 is predominant in dry seeds, which may indicate that it is important for sensing the conditions for germination. On the other hand, Phot-2 is predominant in petals, so it may have a particular role here.
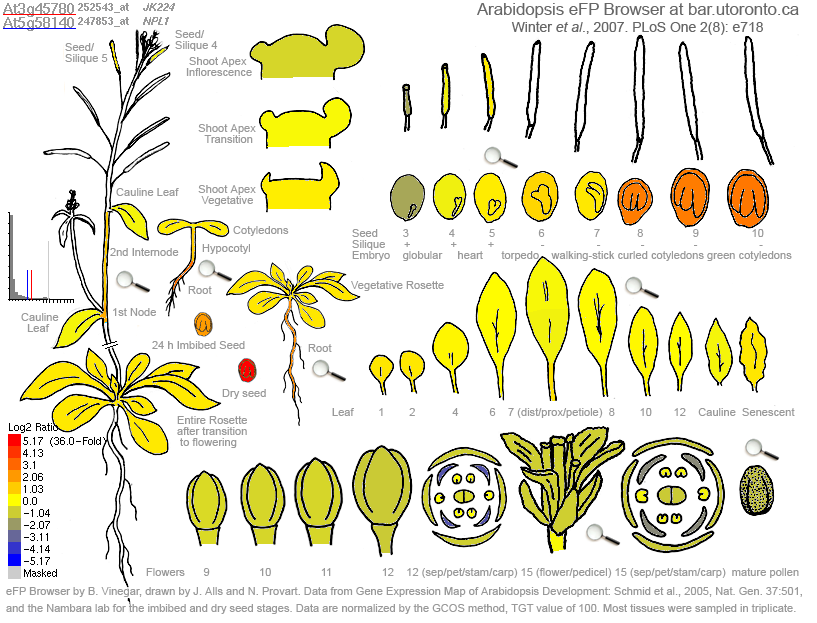
Figure 3. Comparison of mRNA expression between Phot-1 (At3g45780) and Phot-2 (At5g58140). Redder colors indicate a predominance of Phot-1 relative to Phot-2, while bluer colors indicate a predominance of Phot-2 relative to Phot-1. Note that this is a logarithmic scale; a deep red indicates a 36-fold degree of predominance. Yellow indicates an equal proportion in expression. Leaves have a roughly equal expression. Seeds and later embryos have a predominance of Phot-1. Early embryos and floral tissues, especially petals, have a predominance of Phot-2.
Figure 4 below shows the expression levels of Phot-1 and Phot-2 in the bottom of the root. As before, Phot-1 is especially highly expressed in the cortex region of levels 3, 4, and 5. On the other hand, Phot-2 is dominant in the epidermis and root cap. The differences here should be taken with a grain of salt, however, because of the low levels of expression to begin with. These low levels are subject to much error, which is not indicated here. Direct GFP experiments with two colors (e.g. Phot-1 could be labelled red and Phot-2 green) could be done to more accurately determine the relative abundance of these two photoreceptors in the growing region of the root.
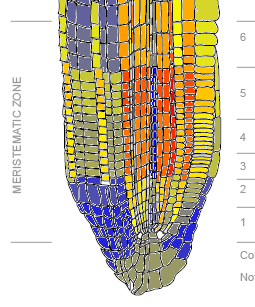
Figure 4. Relative predominance of mRNA expression of Phot-1 (At3g45780) and Phot-2 (At5g58140). Bluer colors indicate a predominance of Phot-2, while redder colors indicate a predominance of Phot-1. Phot-1 is dominant in the cortex and related tissues for levels 3, 4, and 5, while Phot-2 is dominant in the epidermis and root cap.
Summary
Modern bioinformatic technology allows for very large amounts of data to be collected quickly. The standardized nature of gene chips means that mRNA expression levels in different samples can be directly compared. Although it does not directly measure protein abundance as would GFP experiments, it is a good approximation. Repeated gene chip experiments can be used to determine the effect of various treatments on a given protein's expression. In case of the above, repeated gene chip experiments show how Phot-1 and Phot-2 levels vary depending on the cell or tissue type. Interestingly, Phot-1 is localized to a rather specific area of roots. This area may serve as a light-sensitive "eye" that allows for root phototropism.
